
G
H
Hep-2 Cell adherence assay to determine presence of EAEC
Materials
-Dulbecco-MEM supplemented with 10% FBS and 5 ml antibiotic-antimycotic (10,000 units/ml)
-Dulbecco-MEM with 1% D-Mannose, 10% FBS without antibiotics
-Lab-Tek II Chamber Slide w/Cover, 4well
-Tryptic Soy Broth with 1% D-Mannose
Procedure
-Grow Hep-2 cells at 37°C in 5% CO2 in D-MEM supplemented with 10% FBS and 5 ml antibiotic-antimycotic (10,000 units/ml)
-Seed a 4 chamber slide with Hep-2 cells at a concentration 1x105 and allow Hep-2 cells attach to the slide overnight.
-Incubate selected isolates of E.coli overnight at 37°C in Triptic Soy Broth with 1% D-Mannose
-The following morning, remove the cell culture medium in the chamber slide, wash with Hanks’ balance salt solution and replace with D-MEM with 1%
D-Mannose, 10% FBS without antibiotics.
-Add 10µl of E.coli in each well and incubate at 37°C, 5% CO2 for 3 hours.
-After that the slide should be wash vigorously 5 times with PBS
-Fix with 100% Methanol
-Stain with Wright-Giemsa
Materials
-Dulbecco-MEM supplemented with 10% FBS and 5 ml antibiotic-antimycotic (10,000 units/ml)
-Dulbecco-MEM with 1% D-Mannose, 10% FBS without antibiotics
-Lab-Tek II Chamber Slide w/Cover, 4well
-Tryptic Soy Broth with 1% D-Mannose
Procedure
-Grow Hep-2 cells at 37°C in 5% CO2 in D-MEM supplemented with 10% FBS and 5 ml antibiotic-antimycotic (10,000 units/ml)
-Seed a 4 chamber slide with Hep-2 cells at a concentration 1x105 and allow Hep-2 cells attach to the slide overnight.
-Incubate selected isolates of E.coli overnight at 37°C in Triptic Soy Broth with 1% D-Mannose
-The following morning, remove the cell culture medium in the chamber slide, wash with Hanks’ balance salt solution and replace with D-MEM with 1%
D-Mannose, 10% FBS without antibiotics.
-Add 10µl of E.coli in each well and incubate at 37°C, 5% CO2 for 3 hours.
-After that the slide should be wash vigorously 5 times with PBS
-Fix with 100% Methanol
-Stain with Wright-Giemsa
Gel Dry (for
SDS-PAGE gels)
Glycerol (5%) 300 ml
Methanol (10%) 600 ml
ddH2O 5100 ml
Glycerol Frozen
Stock
800 ml of E. coli o/n
culture
Add 200 µl 80% glycerol.
Freeze at -80C or in
liquid nitrogen
(Don't freeze competent
cells in liquid nitrogen;
only at -80°C.
SDS-PAGE gels)
Glycerol (5%) 300 ml
Methanol (10%) 600 ml
ddH2O 5100 ml
Glycerol Frozen
Stock
800 ml of E. coli o/n
culture
Add 200 µl 80% glycerol.
Freeze at -80C or in
liquid nitrogen
(Don't freeze competent
cells in liquid nitrogen;
only at -80°C.

Gel Running Buffer (5X and 1X)
I) Glycine (1.92 M) 576 g
Tris-Base (500 mM) 240 g
QS to 4 L
II) 20% SDS
1X running buffer: 2 L 5X buffer + 50 ml of 20% SDS + 7950 ml H2O.
I) Glycine (1.92 M) 576 g
Tris-Base (500 mM) 240 g
QS to 4 L
II) 20% SDS
1X running buffer: 2 L 5X buffer + 50 ml of 20% SDS + 7950 ml H2O.
Heat Shock (JM100 Competent Cells (Stratagene)
The JM109 strain is appropriate for routine cloning applications. The strain is endocuclease
(enA) deficient, greatly improving the quality of minprep DNA, and is recombination (recA)
deficient, improving insert stability. The hsdR mutation prevents the cleavage of cloned DNA
by the EcoK endocuclease system. The laclqZΔM15 gene, on the F' episome, allows for blue-
white screening.
Protocol:
The JM109 strain is appropriate for routine cloning applications. The strain is endocuclease
(enA) deficient, greatly improving the quality of minprep DNA, and is recombination (recA)
deficient, improving insert stability. The hsdR mutation prevents the cleavage of cloned DNA
by the EcoK endocuclease system. The laclqZΔM15 gene, on the F' episome, allows for blue-
white screening.
Protocol:

Attachment assay using E. coli strain 042.
3X Gel
Buffer
3 M Tris pH
8.45 + 0.3%
SDS
Buffer
3 M Tris pH
8.45 + 0.3%
SDS
The Gram-stain
A sample of a frozen stock colony is grown in LB (or respective media) at 37°C
overnight. The Gram staining can be performed as described by Becton, Dickinson:
Twenty µL of the o/n culture mixture is placed onto a clear glass slide and spread into a
thin, uniform smear. After the smear dries, the slide is heat fixed by a low flame and then
allowed to cool to room temperature. The fixed smear is then immersed in a Coplin
staining jar containing Gram Crystal Violet for one minute. The slide is then dipped in
deionized water, immersed in a Coplin staining jar containing Gram Iodine for one
minute and then rinsed with deionized water. A 3 mL transfer pipette is then used to
wash the slide with Gram Decolorize until the solvent running from the slide is colorless
and the slide is then rinsed with cool deionized water. Next, the slide is immersed in a
Coplin staining jar containing Gram Safranin counterstain for one minute and then
rinsed with deionized water. Slides are allowed to air dry and then examined under oil
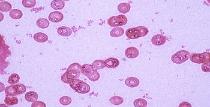
immersion. Isolates that appear blue to purple are considered Gram-positive and those
that appear pink to red are Gram-negative.
GRAM STAIN PRESENTATION
A sample of a frozen stock colony is grown in LB (or respective media) at 37°C
overnight. The Gram staining can be performed as described by Becton, Dickinson:
Twenty µL of the o/n culture mixture is placed onto a clear glass slide and spread into a
thin, uniform smear. After the smear dries, the slide is heat fixed by a low flame and then
allowed to cool to room temperature. The fixed smear is then immersed in a Coplin
staining jar containing Gram Crystal Violet for one minute. The slide is then dipped in
deionized water, immersed in a Coplin staining jar containing Gram Iodine for one
minute and then rinsed with deionized water. A 3 mL transfer pipette is then used to
wash the slide with Gram Decolorize until the solvent running from the slide is colorless
and the slide is then rinsed with cool deionized water. Next, the slide is immersed in a
Coplin staining jar containing Gram Safranin counterstain for one minute and then
rinsed with deionized water. Slides are allowed to air dry and then examined under oil
immersion. Isolates that appear blue to purple are considered Gram-positive and those
that appear pink to red are Gram-negative.
GRAM STAIN PRESENTATION
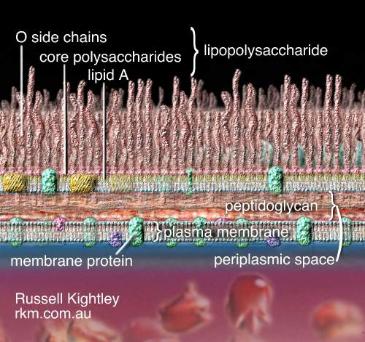
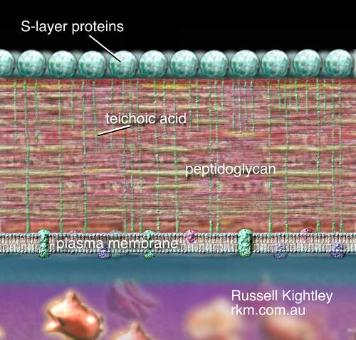
Gram negative Gram positive


S. aureus GENOMIC prep
1. Pellet cells from 10 mls overnight cultures in BHI or TSB.
2. Resuspend pellet in 2 mls of 150 mM NaCl; 10 mM Tris, pH 7.5; 75 mM EGTA pH 7.9.
Transfer the bacterial suspension to glass tubes that can hold ~10 ml and will fit in
your centrifuge.
To make 5 mls: 150 µl 5M NaCl
50 µl 1M Tris pH 7.5
375 µl 1M EDTA
3. Add to the ~2 mls bacterial suspension:
200µl (10 mg/ml) lysozyme
50µl (1 mg/ml) lysostaphin
5µl (10 mg/ml) RNase A
Incubate in the 37°C shaker 1-2h.
4. Add 100 µl of 45% EtOH/SDS saturated solution and incubate at 37˚C.
To make 100 ml: 5g SDS
50 ml 95% EtOH
50 ml dH20 ml
5. Add 650 ul of 5M NaCl and 3 ml of Chloroform/Isoamyl alcohol (24:1). Shake/mix
well (DO NOT VORTEX, will shear the DNA).
6. Centrifuge 12,000 rpm for 15 min.
7. Transfer the clear supernatant to a clean 15 ml conical tube. (DO NOT DISTURB
THE WHITE INTERFACE!).
1. Pellet cells from 10 mls overnight cultures in BHI or TSB.
2. Resuspend pellet in 2 mls of 150 mM NaCl; 10 mM Tris, pH 7.5; 75 mM EGTA pH 7.9.
Transfer the bacterial suspension to glass tubes that can hold ~10 ml and will fit in
your centrifuge.
To make 5 mls: 150 µl 5M NaCl
50 µl 1M Tris pH 7.5
375 µl 1M EDTA
3. Add to the ~2 mls bacterial suspension:
200µl (10 mg/ml) lysozyme
50µl (1 mg/ml) lysostaphin
5µl (10 mg/ml) RNase A
Incubate in the 37°C shaker 1-2h.
4. Add 100 µl of 45% EtOH/SDS saturated solution and incubate at 37˚C.
To make 100 ml: 5g SDS
50 ml 95% EtOH
50 ml dH20 ml
5. Add 650 ul of 5M NaCl and 3 ml of Chloroform/Isoamyl alcohol (24:1). Shake/mix
well (DO NOT VORTEX, will shear the DNA).
6. Centrifuge 12,000 rpm for 15 min.
7. Transfer the clear supernatant to a clean 15 ml conical tube. (DO NOT DISTURB
THE WHITE INTERFACE!).